Distance Metrics: Measuring Similarity
There are multiple ways to measure "similarity" between two embedding vectors. Each has trade-offs. This section covers the math and practical implications.
Cosine Similarity (Most Common)
Definition
Given normalized vectors \(\hat{\vec{u}}\) and \(\hat{\vec{v}}\) (each with magnitude 1):
Equivalent n-dimensional form using summation notation:
This equals the cosine of the angle between vectors:
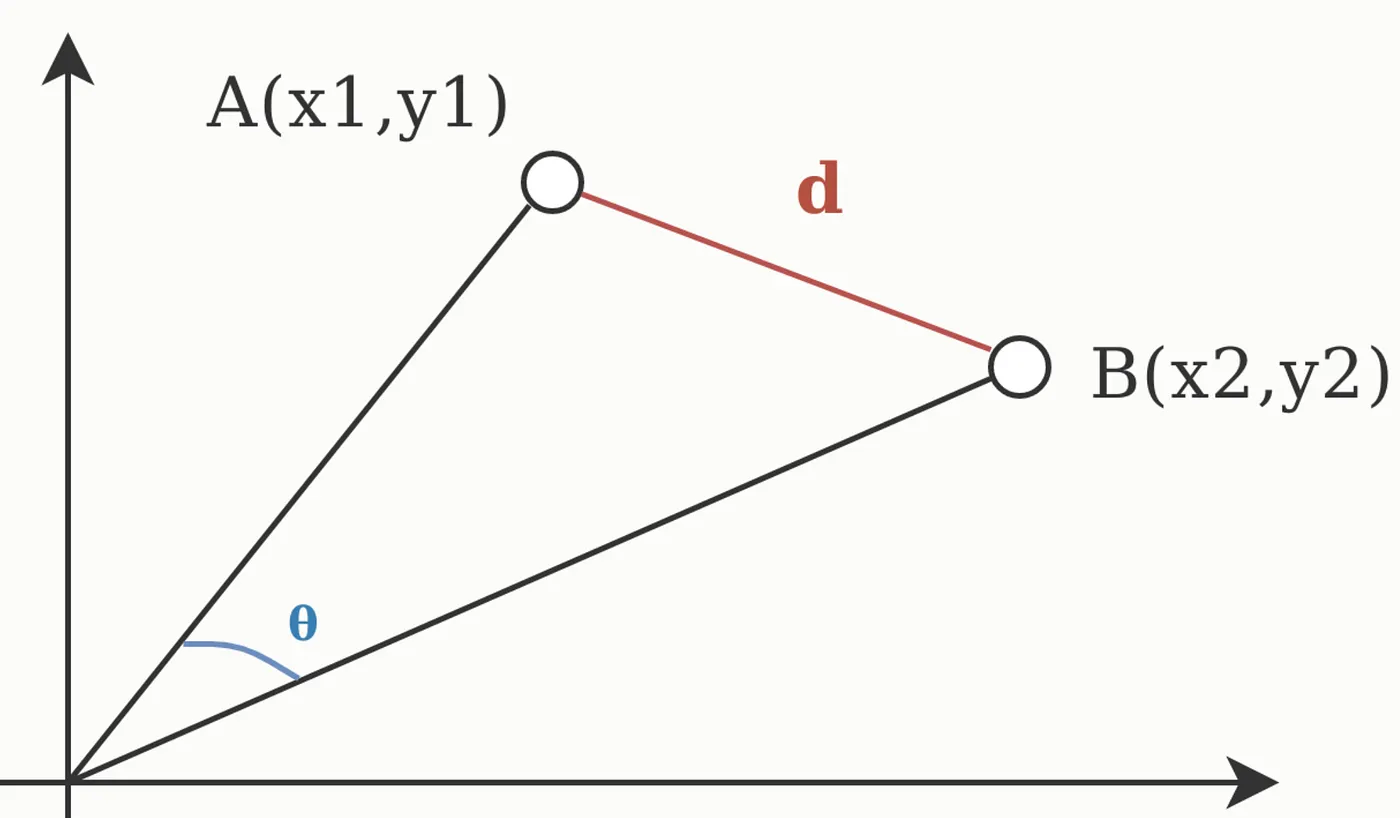
Properties
- Range: \([-1, 1]\)
- 1: identical direction (most similar)
- 0: perpendicular (completely different)
- -1: opposite direction (least similar)
Why It's Used in RAG
- Normalized embeddings: Most models output unit-length vectors
- Angle-based: Captures "direction" not "magnitude"
- Fast: Just a dot product after normalization
- Invariant to scale: Longer passage ≈ shorter passage if similar content
Example
import numpy as np
# Two passages about cats
vec_passage1 = np.array([0.1, 0.9, 0.2, 0.0]) # (normalized)
vec_passage2 = np.array([0.15, 0.85, 0.25, 0.05]) # (similar, normalized)
vec_passage3 = np.array([0.8, 0.1, 0.05, 0.3]) # (about dogs, normalized)
# Cosine similarity
sim_1_2 = np.dot(vec_passage1, vec_passage2)
sim_1_3 = np.dot(vec_passage1, vec_passage3)
print(f"Cat-Cat similarity: {sim_1_2:.3f}") # ≈ 0.975
print(f"Cat-Dog similarity: {sim_1_3:.3f}") # ≈ 0.256
When to Use
✅ Use cosine similarity when:
- Vectors are normalized (most embedding models do this)
- Document length varies widely
- You care about content direction, not magnitude
❌ Avoid when:
- Vectors have important magnitude information
- Document length is meaningful signal
Euclidean Distance
Definition
For vectors \(\vec{u}\) and \(\vec{v}\):
This is the straight-line distance in space.
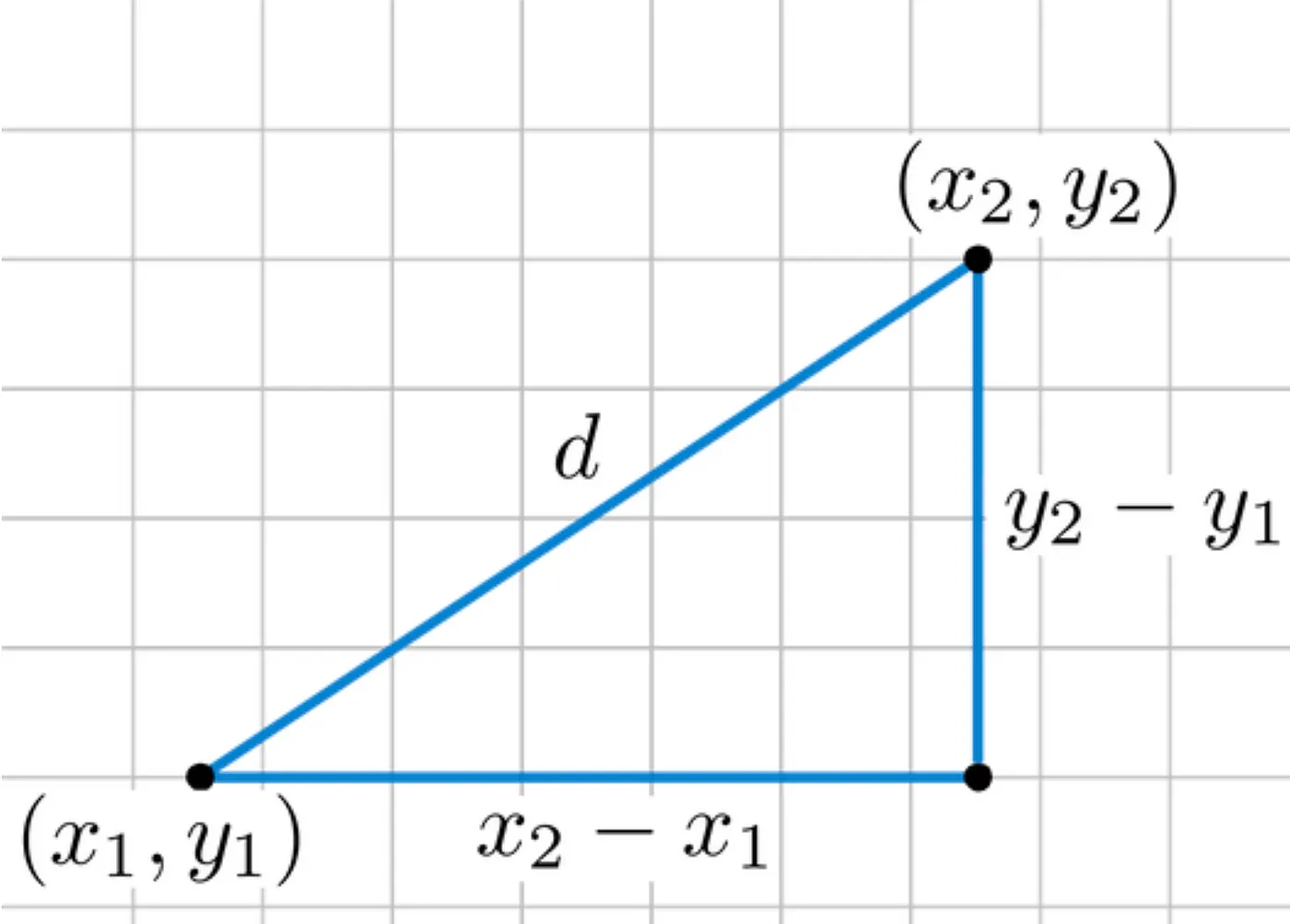
Properties
- Range: \([0, \infty)\)
- 0: identical vectors
- Larger values: more different
- Sensitive to magnitude: longer documents have larger distances
Relationship to Cosine
For normalized vectors, there's a mathematical relationship:
So Euclidean distance and cosine similarity give similar ranking for normalized vectors, but Euclidean is slower to compute.
Example
import numpy as np
u = np.array([1, 2, 3])
v = np.array([4, 5, 6])
# Manual
euclidean = np.sqrt((4-1)**2 + (5-2)**2 + (6-3)**2)
print(f"Euclidean distance: {euclidean:.3f}") # 5.196
# Using numpy
euclidean = np.linalg.norm(u - v)
print(f"Using numpy: {euclidean:.3f}") # 5.196
# Using scipy
from scipy.spatial.distance import euclidean as scipy_euclidean
print(f"Using scipy: {scipy_euclidean(u, v):.3f}") # 5.196
When to Use
✅ Use Euclidean distance when:
- Vectors are not normalized
- Vector magnitude is meaningful
- You're in low dimensions
❌ Avoid in RAG because:
- Embedding models output normalized vectors
- Slower than cosine similarity
- Less semantically meaningful for text
Dot Product (Inner Product)
Definition
Simply the dot product without normalization:
Properties
- Range: \((-\infty, \infty)\)
- Larger values: more similar
- Sensitive to magnitude: longer vectors = larger products
- Fast: single multiplication and sum
Comparison to Cosine
For normalized vectors (\(\|\vec{u}\| = \|\vec{v}\| = 1\)):
But for unnormalized vectors, dot product depends on both direction AND magnitude.
Example
# Unnormalized vectors
u = np.array([2, 3, 4])
v = np.array([1, 2, 3])
# Dot product
dot = np.dot(u, v)
print(f"Dot product: {dot}") # 2*1 + 3*2 + 4*3 = 2 + 6 + 12 = 20
# If we scale u, dot product changes even though direction is same
u_scaled = u * 2
dot_scaled = np.dot(u_scaled, v)
print(f"Dot product (u scaled): {dot_scaled}") # 40
# But cosine similarity stays the same
cos_sim_1 = np.dot(u, v) / (np.linalg.norm(u) * np.linalg.norm(v))
cos_sim_2 = np.dot(u_scaled, v) / (np.linalg.norm(u_scaled) * np.linalg.norm(v))
print(f"Cosine similarities: {cos_sim_1:.3f}, {cos_sim_2:.3f}") # Same!
When to Use
✅ Use dot product when:
- Vectors are normalized (same as cosine then)
- You need maximum speed
- Magnitude information is important
❌ Avoid when:
- Vectors have varying magnitudes
- You need scale-invariant comparison
Manhattan Distance (L1)
Definition
Like the distance traveled on a city grid (you can only move horizontally/vertically).
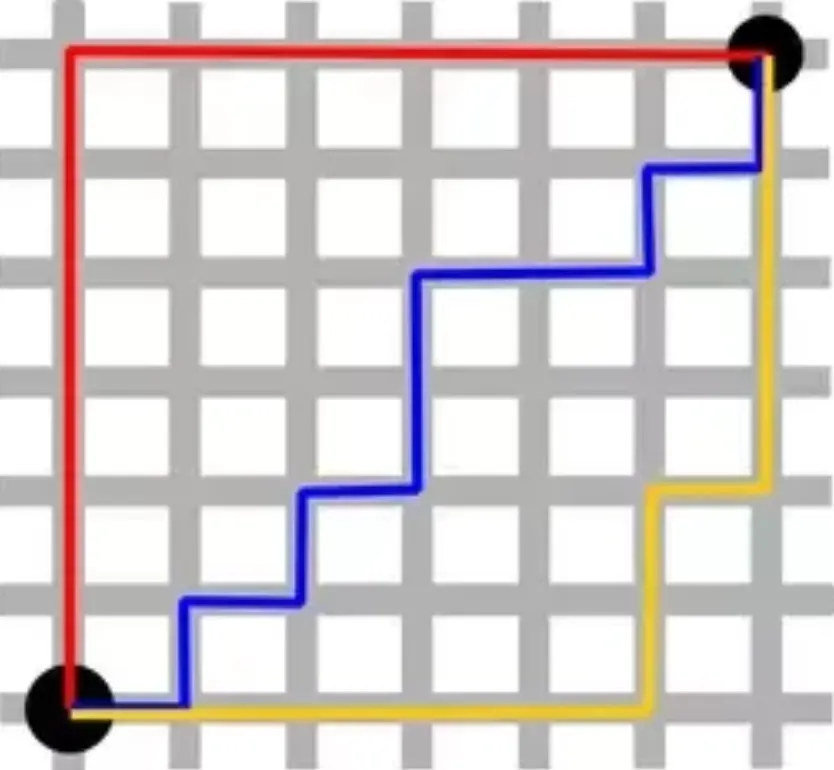
When to Use
Rarely in RAG, but sometimes useful for: - Sparse vectors (mostly zeros) - Special structured data - More robust to outliers than Euclidean
from scipy.spatial.distance import cityblock
u = np.array([1, 2, 3])
v = np.array([4, 5, 6])
manhattan = cityblock(u, v)
print(f"Manhattan distance: {manhattan}") # |4-1| + |5-2| + |6-3| = 3+3+3 = 9
Comparison Table
| Metric | Formula | Range | Type | RAG Use | Speed |
|---|---|---|---|---|---|
| Cosine | \(\frac{\vec{u} \cdot \vec{v}}{\|\vec{u}\|\|\vec{v}\|}\) | [-1, 1] | Angle | ✅ Best | Fast |
| Euclidean | \(\sqrt{\sum(u_i-v_i)^2}\) | [0, ∞) | Distance | ⚠ Avoid | Slow |
| Dot | \(\sum u_i v_i\) | (-∞, ∞) | Similarity | ✅ Good | Fast |
| Manhattan | \(\sum \|u_i-v_i\|\) | [0, ∞) | Distance | ❌ Rare | Medium |
Which Metric to Choose?
Decision Tree:
Are your vectors normalized to unit length?
├─ YES → Use cosine similarity or dot product
│ ├─ Need maximum speed? → Dot product
│ └─ Standard choice → Cosine similarity
│
└─ NO → Use Euclidean distance
(but consider normalizing first)
For RAG systems:
- 99% of the time: Cosine Similarity
- Some vector DB defaults: Dot Product (equivalent for normalized)
- Rarely: Euclidean or Manhattan
Maximum Inner Product Search (MIPS)
When using dot product, the problem becomes: find vectors with maximum dot product (not minimum distance).
This is called MIPS (Maximum Inner Product Search).
If you normalize vectors to unit length:
So MIPS on normalized vectors = nearest neighbor in cosine similarity.
Code Example with Different Metrics
from sentence_transformers import SentenceTransformer
import numpy as np
from sklearn.metrics.pairwise import cosine_similarity
model = SentenceTransformer('all-MiniLM-L6-v2')
# Create some embeddings
docs = [
"Cats are cute and furry",
"Dogs are loyal pets",
"Cats like to sleep and play",
"Python is a programming language"
]
embeddings = model.encode(docs)
# Query
query = "I like my cat"
query_emb = model.encode([query])[0]
# Different metrics
cosine_sims = cosine_similarity([query_emb], embeddings)[0]
dot_products = np.dot(embeddings, query_emb)
euclidean_dists = np.linalg.norm(embeddings - query_emb, axis=1)
# Rank by each metric
cosine_rank = np.argsort(-cosine_sims)[:3]
dot_rank = np.argsort(-dot_products)[:3]
euclidean_rank = np.argsort(euclidean_dists)[:3]
print("Top 3 by cosine:", [docs[i] for i in cosine_rank])
print("Top 3 by dot:", [docs[i] for i in dot_rank])
print("Top 3 by euclidean:", [docs[i] for i in euclidean_rank])
# For normalized embeddings, cosine and dot give same ranking
print("Cosine and dot rankings same?", np.array_equal(cosine_rank, dot_rank))
Summary
| Concept | Use In RAG |
|---|---|
| Cosine similarity | ✅ Best choice for RAG |
| Dot product | ✅ Good if normalized |
| Euclidean | ⚠ Avoid (slower, less meaningful) |
| Manhattan | ❌ Rarely useful |
Next Steps
→ Exact vs Approximate Search - How to search millions of vectors fast
→ Vector Databases - Production systems